On Friday, Jan. 23, I had the privilege of attending the University of Maryland’s Spatial Biology Symposium, which featured talks on developmental biology, cancer and neuroscience. In this piece, I highlight one particular talk that caught my interest — given by Elana Fertig, Dean E. Albert Reece Endowed Professor of Medicine at the University of Maryland. Fertig’s talk centered around rethinking how we can predict and monitor the carcinogenesis of pancreatic cancer through a spatial biology and computational lens. Although my knowledge in the field of spatial biology is limited, frankly, I would like to offer my best understanding of the key ideas and takeaways.
“The [pancreatic] tumor has a very dense, immunosuppressive microenvironment and is basically universally resistant to almost any therapeutic that's given to it,” Fertig explained. “The vast majority of the tumor is macrophages, fibroblasts and other components of the microenvironment. So it makes it really critical, as we start to understand this tumor, to take a spatial biology lens in order to figure out what's in there, what's going on and how all these different cell types interact.”
Fundamentally, spatial biology involves analyzing which cells are present in a specific tissue, where they are distributed and how they interact with each other to shape the tissue microenvironment. Because of the complexity of tissue biology, answering these questions requires a combination of powerful tools like proteomics and transcriptomics, which together provide complementary information about gene expression, protein abundance and signaling activity within individual cells and a collective tissue. It turns out this information can help us understand a lot about cells.
“When we have these high-dimensional assays, we can go beyond what cell types are there. Where's the cell phenotypically? Where is it temporally? In terms of evolution? Where is it in the cell cycle? Where is it spatially?” Fertig said.
Indeed, high-dimensional assays allow researchers to move beyond simply identifying cell types and allow them to uncover the biological programs operating within those cells. A pancreatic tumor cell, for instance, may be proliferating, responding to inflammatory signals, transitioning into a more invasive state or interacting dynamically with neighboring fibroblasts or immune cells. Each of these processes leaves a distinct molecular footprint in transcriptomic and proteomic data. Fertig’s team uses computational models to interpret and organize these complex molecular signals.
Interestingly, the mathematical concept of matrix factorization is especially useful to decoding biological patterns.
“The axes of identity of different cellular signatures perfectly map to the mathematics of matrix factorization,” Fertig said. “Matrix factorization lets you learn patterns in the data, learn what are the features that are being used in different cellular groups… and what are the genes that are being associated with each pattern.”
Using matrix factorization, the team uncovered distinct cellular programs and identified a significant association between epithelial-to-mesenchymal transition (EMT) signaling and cancer-associated fibroblasts. It thus becomes clear that mathematical approaches often help researchers model biological systems at large. In fact, this kind of mathematical modeling is highly prevalent in the field of computational biology.
Fertig highlighted the driving question of such mechanistic mathematical models: “If I know what cells or molecules do, can I build up a system of equations from first principles and use those to probe the system?”
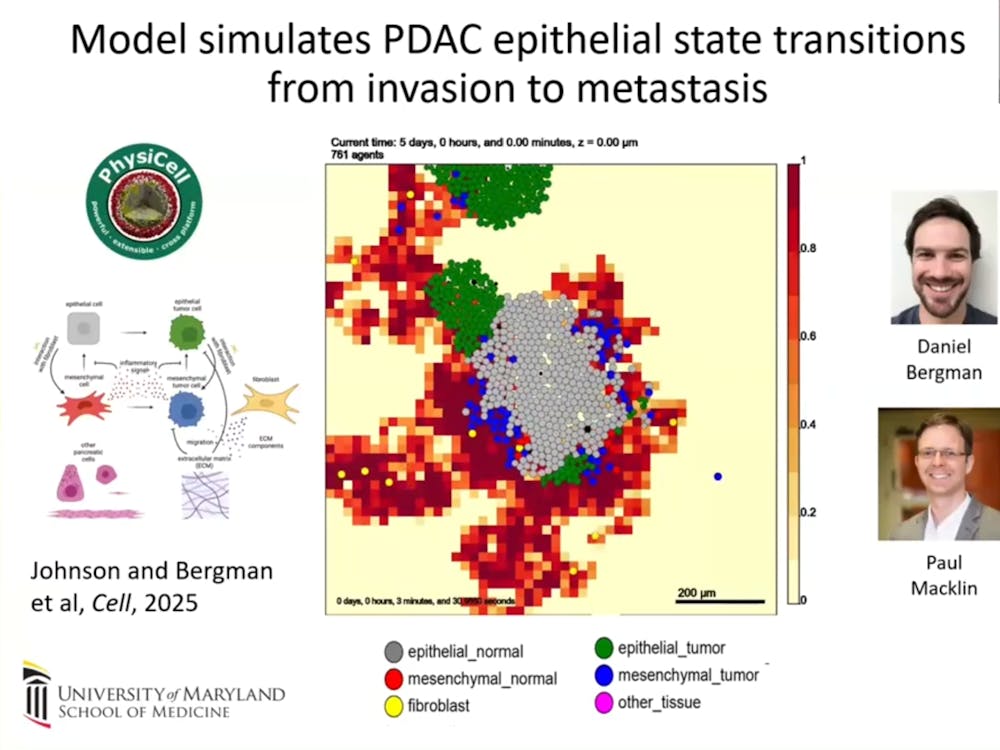
In other words, mechanistic models can move beyond simply analyzing data to simulate biological systems by actually encoding the fundamental rules that govern cellular behavior. In fact, Fertig and her collaborators used this approach to apply PhysiCell, a virtual platform that simulates how cells grow, evolve and interact with their environment over time to pancreatic cancer. The simulation revealed how normal epithelial cells transition to mesenchymal based on their interactions (EMT), which is hypothesized to be a major mechanism for the proliferation of pancreatic cancer.
“The [epithelial] cells start to transition when they interact with fibroblasts. After some time, the EMT-like cells are moving a little bit more and are able to go out and get away from the fibroblasts and escape. Eventually, when the cells get far enough away, they're able to colonize and start seeding out tumors,” Fertig explained, referencing the simulation.
Not only can mathematical-based simulations illuminate such obscure mechanisms, but they are also convenient: researchers can simply adjust the parameters and perform as many experiments as needed on their computer. One of the main challenges, however, lies in determining the mathematical rules in the first place. Fortunately, powerful tools such as transcriptomics in combination with bioinformatics software can help with this.
“Let's say you have your single-cell or spatial data set. Now, that [bioinformatics] software will let you read out the labels of your single-cell object in the metadata and automatically pull down what cell types are there, so that you can start to form the rules that you would want to encode,” Fertig clarified. “In the case of spatial transcriptomics, we can initialize the cells in the right density in different types of spatial distributions.”
In spite of the power and utility of these mathematics-based computational models, it is important to ensure that the findings actually reflect biological reality and are not just “computational artifacts.” To that end, researchers turn to good old, hands-on experimentation to corroborate their results.
Nevertheless, the times are certainly changing. As science at large, not just spatial biology, shifts from purely data-driven to computational tools and modeling, we should remain critical instead of blindly following the trends. How can we use these approaches to refine our hypotheses? To ask essential, deeper questions? To refine our experiments? For me, hearing what experts have to say in talks like this reminds me of how much there still is to learn.