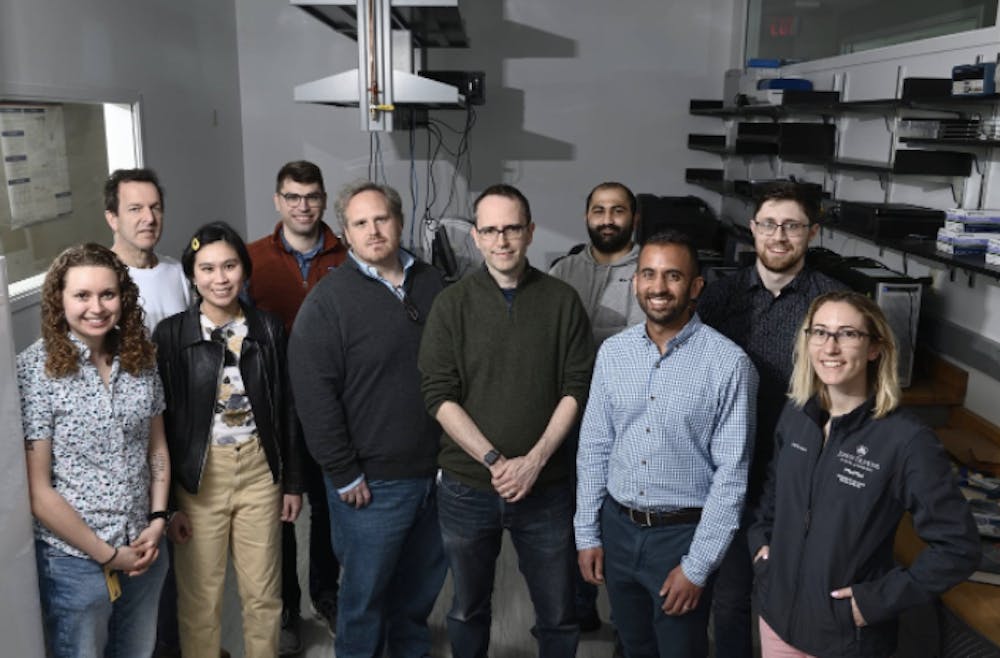
Researchers from Hopkins have contributed to a group that has analyzed the first complete sequence of a human genome, led by Rajiv McCoy, Michael Schatz and Winston Timp. Their work is part of the Telomere-to-Telomere (T2T) consortium of over 100 researchers globally.
McCoy, an assistant professor in the Department of Biology, discussed their research in an interview with The News-Letter.
“It’s the biggest project I’ve ever been a part of. It was amazingly collaborative and grassroots — a very inclusive group,” he said.
This discovery builds on the work of the Human Genome Project and will allow the scientific community to better understand how DNA affects disease risk and gene expression. The Human Genome Project, an international research project completed in 2003, established a reference genome of human DNA, which is a template used as a standard of comparison in research.
The McCoy lab, which contributed to the T2T project, focuses on population genetics and evolutionary aspects of genomic analysis.
“[The genome] contains the basic instructions for human life, and it’s a remnant of our evolution,” McCoy said. “Looking at patterns of variation across genomes can tell us about our history — where we come from. Having a complete view of it is something that’s really fundamental to our understanding of human biology.”
He explained that the Human Genome Project succeeded in analyzing most of the human genome, but due to technical limitations of the time, some sequences were left unresolved.
“Almost all genomic analyses rely on taking sequencing rates from a given sample and comparing them back to that reference genome,” McCoy said. “Even the current, latest version of that genome is still missing about 8% of really challenging sequences.”
McCoy explained that technical improvements in sequencing platforms have made it possible to identify the missing portions of the reference genome.
“You can think of this process of genome assembly sort of like putting together a puzzle,” he said. “If you have a puzzle with a lot of pieces, and there are parts of the puzzle where there’s a lot of one color, for example, it can be really hard to figure out how to piece those together.”
The development of long-read technology was critical to the sequencing of the complete human genome. Long-read sequencing allows longer strings of DNA base pairs to be read at once. This allows more variants to be detected since more base pairs are being analyzed at once.
“Long reads are like having much bigger puzzle pieces, so there’s a greater likelihood that you’ll catch something in one of those big pieces that allow you to place that puzzle piece in the context of the rest of the puzzle,” McCoy said. “For these genome assemblies, you can span some of those very repetitive regions and place those sequences with respect to the rest of the genome.”
Six papers from the project have been published in a special issue of Science. The first paper introduces the completed human genome, while the other five papers explain how this discovery will improve our understanding of human biology.
This complete reference genome includes 200 million base pairs of sequences that have never been resolved before. It uses DNA from a hydatidiform mole and represents the genome of a single individual, while the original reference genome is a composite of about 20 individual genomes, McCoy explained.
In an interview with The News-Letter, Stephanie Yan, a co-first author on one of the papers and PhD biology student at the Krieger School of Arts and Sciences, explained her goals for the project.
“I’m hoping it’s the very first step in a different way of approaching how people in human genetics do genetic studies at all,” Yan said.
The McCoy lab is continuing research in this field by analyzing additional functional genomic data with respect to the new reference genome.
Melanie Kirsche, a PhD computational genomics student at the Whiting School of Engineering, is another co-first author on the project. She studied how the new genome sequence can improve genomic analyses, including aligning reads from other individuals with it, and how the genome varies between individuals.
In an interview with The News-Letter, Kirsche expressed hope that in the future, the field will work toward making genomic sequencing more automated, allowing researchers to sequence many other human genomes with diverse ancestry.
The researchers involved each expressed gratitude for being able to contribute to this project.
Yan explained that she heard about the T2T project in summer 2020, five months before she became directly involved in it.
“It was such a huge honor for me when Mike [Schatz] initially invited me to join the project. I was like, ‘I get to work on T2T? I get to work on this awesome project that I’ve heard so much about and that I think is a huge milestone for the field of genetics?’ That was just so exciting,” Yan said.
For Kirsche, the T2T research made up a large portion of her PhD thesis.
“It was just a very exciting project to work on for me, to be able to collaborate with probably around 100 researchers from all around the world who are some of the experts in genome assembly and various aspects of genomics,” Kirsche said. “This is the highest-impact thing that I’ve worked on in my PhD, because it’s probably going to be used for years and years to come.”
Similarly, McCoy found it rewarding to complete the work begun by the Human Genome Project.
“The Human Genome Project’s first announcement was when I was in middle school — I was following it on the news,” McCoy said. “I had no idea that I’d be able to participate in this next step in the project and to have my lab participate in this as well.”