In the field of microbial ecology, a positive virus test isn’t always a bad thing. Of course, the viruses in most ecological studies aren’t the kind infecting humans or making headlines every night. Rather, scientists like Eric Sakowski are interested in the distribution and impact of bacteriophages, or viruses that infect bacteria.
At times, infection can improve the health of an ecosystem For example, bacteriophages could prevent the overgrowth of cyanobacteria, causing algal blooms. On the other hand, if the bacterial population decreases too low, the effects could be disastrous. Like porridge in “Goldilocks,” viral infections in bacterial populations have to be just right.
Sakowski is a former postdoctoral researcher at Hopkins and a current professor at Mount St. Mary’s University. Sakowski, who has always been interested in the role of viruses in aquatic ecosystems, worked in the lab of Environmental Health and Engineering Professor Sarah Preheim.
The two both acknowledged that there was not a good technique for detecting viral infections in microbial ecosystems, and so Sakowski combined his interest in virology with Preheim’s knowledge of a technique known as emulsion, paired isolation and concatenation polymerase chain reaction (epicPCR).
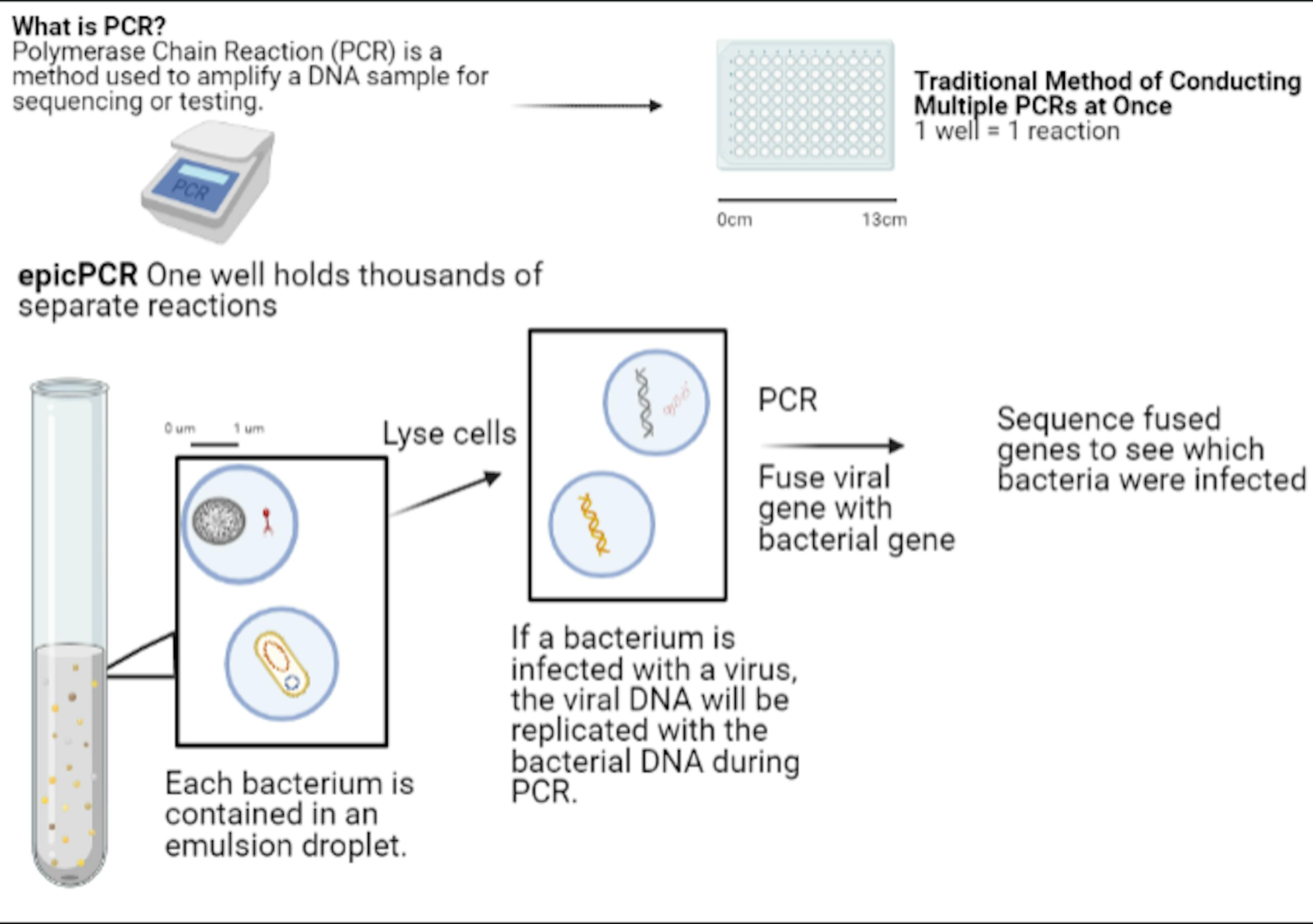
Traditional polymerase chain reaction (PCR) helps to multiply a DNA sample until there is enough of it for analysis. The technique is used in forensic science, biology research and even COVID-19 tests. However, the process becomes rather limited and tedious when one is seeking to analyze the DNA of several different cells, such as the bacteria in a waterway. According to Preheim, that’s where epicPCR comes in.
COURTESY OF ELLIE ROSE MATTOON / FIGURE CREATED IN BIORENDER.COM
The bubbles formed by epicPCR can be compared to small test tubes.
“Emulsion just kind of works like an individual test tube,” Preheim said in an interview with The News-Letter.
Each cell is trapped in a bubble-like emulsion, and an infected bacterial cell becomes trapped with the offending virus. Once together, the viral and bacterial genomes fuse, allowing for researchers to see which virus infected what bacteria based off sequencing data. While a traditional “well plate” method pictured below could test a few hundred cells, Sakowski stated that epicPCR could do much more.
“We can get thousands upon thousands of these individual droplets, amounting to tens of thousands of cells at one time,” Sakowski said.
Preheim was involved in the initial development of epicPCR as a postdoctoral researcher at the Massachusetts Institute of Technology. While the technique has been used to detect nitrogen or sulfur cycling genes in a bacterial population, this is the first time it has been used successfully to detect viral genes.
In 2015, Sakowski and Preheim first set out to accomplish this feat, but the process was not easy.
“It was about two years of failure and really just troubleshooting the method,” Sakowski said.
While the original method of epicPCR was complicated and involved toxic reagents, Sakowski was willing to put in the extra time to ensure the method was safer, easier and accessible to any laboratory. Once the technique was developed, the group turned to the Chesapeake Bay for testing.
When analyzing the results, Preheim and Sakowski agreed that because so much about the field was unknown before this technique was developed, questions abound more often than answers.
Sakowski mentioned an interest in further investigating their finding on the impact of tides on phage infections. At high tide and low tide, the researchers noted the same bacteria but different viruses. In addition, the group wants to capture the interactions of cyanobacteria with viruses, as these are an important set of bacteria for environmental health.
At the same time, epicPCR is still a new technique, and work still needs to be done to perfect it.
“You’re always trying to continue improving the method because then it allows you to keep asking better and better questions about the actual ecology,” Sakowski said.
Eventually the method could even be developed for examining the health of the human gut microbiome.
Preheim expressed the sentiment that microbial health is an important question to continue examining, as bacteria are known to play a crucial role in remediating toxic pollution and carrying out biogeochemical cycles.
“I could go on for a long time about all of the wonderful things microbes can do,” Preheim said.
For all the bacteria on the planet, there are 10 times as many viruses. Their role in the ecosystem is less well-known, but hopefully with the advent of new techniques, the scientific community will be able to better determine the role they play in sustaining life as we know it.